[ad_1]
The blood cancer had returned, and Kevin Sander was running out of treatment options. A stem-cell transplant would offer the best chance for long-term survival, but to qualify for the procedure he would first need to reduce the extent of his tumour — a seemingly insurmountable goal, because successive treatments had all failed to keep the disease in check.
As a last throw of the dice, he joined a landmark clinical trial. Led by haematologist Philipp Staber at the Medical University of Vienna, the study is exploring an innovative treatment strategy in which drugs are tested on the patient’s own cancer cells, cultured outside the body.
In February 2022, researchers tried 130 compounds on cells grown from Sander’s cancer — essentially trying everything at their disposal to see what might work.
One option looked promising. It was a type of kinase inhibitor that is approved to treat thyroid cancer, but it is seldom, if ever, used for the rare subtype of lymphoma that Sander had. Physicians prescribed him a treatment regimen that included the drug, and it worked. The cancer receded, enabling him to undergo the stem-cell transplant. He has been in remission ever since. “I’m a bit more free now,” says Sander, a 38-year-old procurement manager living in Podersdorf am See, Austria. ”I do not fear death any more,” he adds. “I try to enjoy my life.”
His story is a testament to this kind of intensive and highly personalized drug-screening method, referred to as functional precision medicine. Like all precision medicine, it aims to match treatments to patients, but it differs from the genomics-guided paradigm that has come to dominate the field. Instead of relying on genetic data and the best available understanding of tumour biology to select a treatment, clinicians throw everything they’ve got at cancer cells in the laboratory and see what sticks.

Is precision public health the future — or a contradiction?
But what it sometimes lacks in elegance, it could make up for in results: in pilot studies, Staber and his colleagues found that more than half of people with blood cancer whose treatment was guided by functional drug testing enjoyed longer periods of remission compared with their experiences of standard treatments1,2. Large-scale testing of genome-directed approaches suggests that the techniques are very effective against some cancers, yet they benefit, at most, only around 10% of patients overall3. Staber and his group’s latest trial is the first to compare functional- and genome-guided approaches head-to-head alongside treatments directed by standard pathology and physician intuition.
“That’ll be a very powerful study, and it will probably vindicate the utility of these functional assays,” says Anthony Letai, a haematologist at the Dana-Farber Cancer Institute in Boston, Massachusetts, and president of the Society for Functional Precision Medicine, a professional organization founded in 2017 to advance the field. And, if anecdotal reports serve as any indication, the try-everything tactic seems to bring about meaningful improvements, even when the genetic sequence of a tumour provides no actionable information, as was the case for Sander.
Companies around the world are already offering these kinds of personalized drug testing service. But proponents of the strategy still have much to prove. Although the concept of screening a bunch of drugs seems simple, the methods used to culture cancer cells outside the body can be technically demanding, time-consuming and costly.
The challenges are particularly acute for solid tumours, which live in complex environments inside the body; replicating those conditions is no easy feat. Researchers are trying wildly differing methods that range from growing tumour samples in mice and chicken embryos to cultivating carefully engineered organoids, and even the delivering infinitesimal amounts of various medicines to a tumour while it’s still in a patient.
Figuring out what works and what is practical, with regard to cost and scale, won’t be easy. But momentum is growing, says Christopher Kemp, a cancer biologist at the Fred Hutchinson Cancer Center in Seattle, Washington. “This is a revolution. Patients are demanding this approach.”
Behind the screen
Down a long corridor, beyond a set of tangerine-coloured doors, lies the Vivi-Bank at the Medical University of Vienna. Short for ‘Viable Biobank’, the room is brimming with liquid-nitrogen dewars, each containing frozen lymphoma samples.
When surgeons extract biopsies from cancerous lymph nodes, they usually immerse the tissue in formaldehyde to prepare for standard pathology analyses. That kills the cells, however, rendering them useless for functional testing. So, to enable drug screens, Staber and haematopathologist Ingrid Simonitsch-Klupp, who jointly oversee the Vivi-Bank, had to convince their surgical colleagues to change their ways, keeping the tissue alive and sending it quickly for processing and storage. “Fresh tissue is the most important thing,” Simonitsch-Klupp says.
Some of that tissue arrives in Staber’s lab, where researchers break up the cells using a knife, forceps and a nylon strainer, creating a slurry to distribute across a 386-well plate. In each well, they test a different drug compound — chemotherapy agents, enzyme-targeted drugs, immune-modulating therapies and more. After a night of incubation, lab testing reveals which drugs are active against the cancer and which ones are not.
A team of clinicians, known as a molecular tumour board, then uses this information to determine the most appropriate course of treatment for each patient.
Several groups have reported success with this general approach. In a trial from the University of Helsinki, for example, researchers found that individualized drug screening of leukaemia cells provided informative results substantially faster than did genomic profiling, yielding impressive clinical responses as well4. Of 29 people with treatment-resistant acute myeloid leukaemia (AML), 17 responded to drug-screening-informed therapies and entered remission.
Likewise, Candace Howard, a radiologist at the University of Mississippi Medical Center in Jackson, and her colleagues published a study last year showing that people with aggressive brain tumours live longer when their chemotherapy regimens are guided by lab testing than when their treatment is directed by a physician’s intuition alone5 — with lower annual health-care costs to boot6.
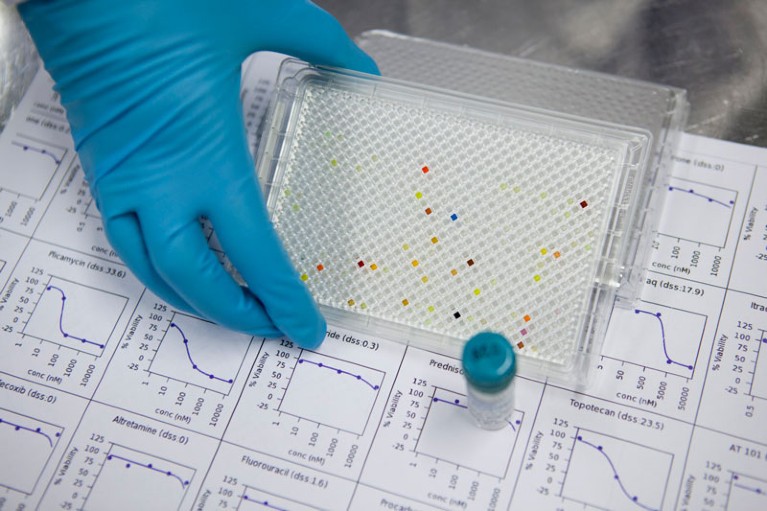

Multi-well plates can be used to test the effectiveness of many cancer drugs at once.Credit: FIMM, University of Helsinki
“It’s cheaper and it’s more effective,” says Jagan Valluri, a cell biologist at Marshall University in Huntington, West Virginia, who co-founded a company called Cordgenics, also based in Huntington, to commercialize the assay used in Howard’s trial.
Functional drug testing is not a new idea. It was embraced by cancer researchers in the late twentieth century, but soon fell out of favour — largely owing to the limitations of assays at the time and a restricted repertoire of anti-cancer drugs. Technological improvements and an expanded pharmacopoeia have changed the picture. Yet, as with most lab-based testing systems, the necessary equipment can be expensive and requires trained personnel to operate it.
That’s a big limitation according to Joan Montero, a biochemist at the University of Barcelona in Spain, because it hinders the broad implementation of functional precision drug testing, especially in low-resource settings. To address these challenges, Montero and his colleagues have been developing inexpensive and portable microfluidic devices for rapid, on-site testing of cancer cells7.
Their microfluidic platform remains years away from practical use, however. And it might guide treatment only for certain types of cancer. That’s because protocols developed for tailoring therapies against blood cancers do not always work in solid tumours of the breast, lung, liver and other organ systems.
Biopsies from solid tumours often yield lower cell counts, requiring extra steps to culture the cells before drug screening. Moreover, solid tumours have complex interactions with healthy cells in their surroundings, meaning that models should be more sophisticated.
Growing pains
The first challenge remains growing enough tumour tissue to test. David Ziegler, a paediatric neuro-oncologist at Sydney Children’s Hospital in Australia, had set out to perform individualized drug screens for around 1,000 children with high-risk cancers as part of the country’s Zero Childhood Cancer Program. But in pilot testing, he and his team discovered that, after several days under lab conditions, up to one-fifth of the patient samples either contained no cancer cells at all, or the cancer cells were being outcompeted by normal, healthy cells8. The researchers quickly learnt to check cultures for tumour cells — through imaging, cellular analysis or genetic profiling — before testing them against drugs.

Super-precise CRISPR tool enters US clinical trials for the first time
Cell cultures from solid tumours can, in principle, be subjected to the same kind of testing used for blood cancers. But an increasing number of research teams are crafting elaborate structures, known as organoids, to test. These patient-derived 3D tissue models — made by growing tumour samples in specialized scaffolds over the course of several weeks — are designed to replicate the intricate tissue architecture of a tumour, thereby offering a more accurate representation of the cancer that physicians are looking to treat.
“We want to put the tumour cells in an environment that’s as close [as possible] to how they were growing in the body,” says Alice Soragni, a cancer biologist at the University of California, Los Angeles.
The process can add weeks to the timeline for obtaining drug sensitivity data. But the extra effort and time investment is worth it, says Carla Grandori, co-founder and chief executive of SEngine Precision Medicine in Bothell, Washington.
In clinical validation studies, Grandori and her SEngine colleagues found that the drug-screening results using organoids aligned with patient outcomes with around 80% accuracy. Those findings are not yet published, but the company — which counts Kemp among its founders — has put out case reports over the past year describing people with difficult-to-treat cancers who, after seemingly running out of treatment options, found unexpectedly effective remedies through organoid drug testing9,10.
Heidi Gray, a gynaecological oncologist at the University of Washington Medical Center in Seattle, treated one of these patients, a woman with ovarian cancer. “Her response was definitely one of the best I’ve seen,” she says. The drug they tried is generally used to treat leukaemia, but it helped to beat back the woman’s ovarian tumour for more than a year, allowing her to travel and enjoy precious time with loved ones before ultimately succumbing to the disease. “We profoundly improved her quality of life,” Gray says, “and that would not have happened without the knowledge provided by this test.”
Model of efficiency
In the hope of testing drugs against even more realistic cancer systems, some researchers have opted to study mice implanted with fresh tumour specimens, a model system known as a patient-derived xenograft.
These personalized ‘avatars’ were once heralded as the next big thing in cancer care. But it soon became evident that many tumours do not grow in mice, that drug screening in xenografts takes too long to provide timely recommendations and that the cost of the approach — often exceeding US$50,000 — is more than most patients and health-care systems can bear.
“It was too slow, too expensive and not robust enough,” says David Sidransky, an oncologist at Johns Hopkins University School of Medicine in Baltimore, Maryland, and a co-founder of Champions Oncology, a leading developer of xenograft models, based in Hackensack, New Jersey.
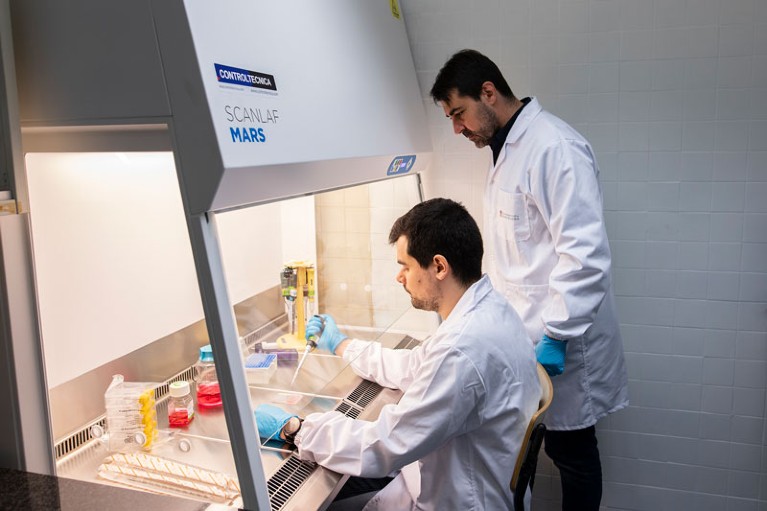

Joan Montero (standing) and his colleagues are developing a low-cost microfluidic device.Credit: University of Barcelona
Although some drug companies continue to use xenografts for research, and some oncologists think that there are certain situations in which they can inform patient care, for the most part, researchers have moved away from mice for functional testing in the clinic. Some have moved on to other living systems.
One such alternative comes from cancer biologist Hon Leong and his colleagues at Sunnybrook Hospital in Toronto, Canada, who devised a system for screening drugs on tumour biopsy samples cultivated on developing chicken embryos. The approach is both rapid and inexpensive, says Leong, allowing researchers to assess different drug options in a matter of weeks rather than the months required for mice.
In ongoing trials focused on advanced breast and kidney cancers that have spread to other parts of the body, Leong and his team have successfully used the chicken-embryo system to identify individuals who would benefit from immune therapies. These are among the most effective cancer treatments today, and a drug class that few other avatar systems can accurately assess, says Leong.
CRISPR cancer trial success paves the way for personalized treatments
Another approach comes from Ross Cagan, a developmental biologist at the University of Glasgow, UK, who uses genomic sequencing and genetic engineering to recreate the unique characteristics of a patient’s tumour in a custom-made fruit fly. This involves introducing mutated forms of cancer-promoting genes or incorporating sequences that restrict cancer-suppressing genes — generally between 5 and 16 alterations in total. Feeding the flies with food containing various medications can then reveal therapeutic regimens that suppress cancer growth, either by acting directly on tumour cells or by influencing the animal’s biology in ways that indirectly impede cancer progression.
This is how Cagan and his colleagues identified a new three-drug cocktail — consisting of a lymphoma treatment, a blood-pressure medicine and an arthritis therapy — that, when administered to a man with a rare tumour of the salivary glands, helped to stabilize the cancer for a year11. In another case, involving a man with an aggressive form of colon cancer, the use of fly avatars guided the team to administer a melanoma drug alongside a bone-strengthening agent, resulting in notable tumour shrinkage and a clinical response that lasted for nearly a year12. A biotech start-up in London called Vivan Therapeutics now offers this bespoke fly-making and drug-screening service for $15,000 per patient.
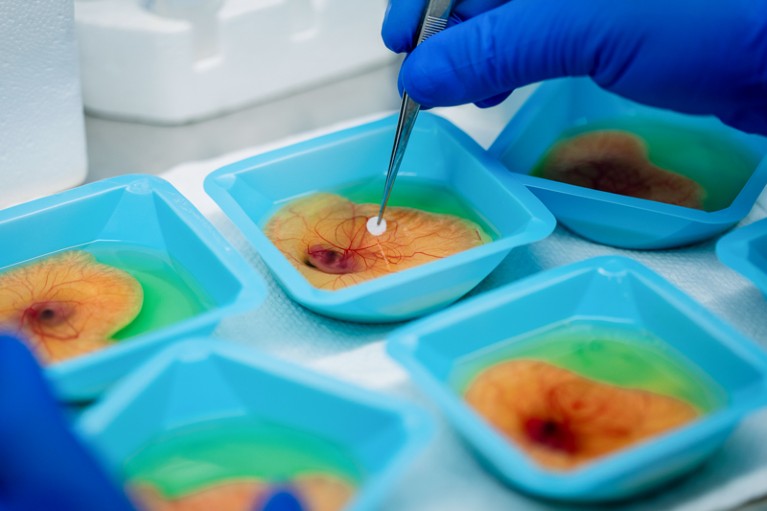
A researcher prepares developing chicken embryos to grow model tumours.Credit: Hon Sing Leong
Any model invariably has biological limitations, however, and so some researchers have elected to do away with animal stand-ins or cellular replicas entirely. Instead, they have developed implantable devices that allow clinicians to test drugs directly on patient tumours — and to do so while the cancer is still inside the body.
Last year, bioengineer Oliver Jonas at Brigham and Women’s Hospital in Boston, and his colleagues demonstrated the feasibility of this strategy in people with lung13 and brain14 cancers. In small trials, surgeons inserted tiny drug-releasing devices, each loaded with nanodoses of up to 12 drugs, into tumours as people underwent cancer-removal surgery. Over the course of the operation, drugs flowed into the surrounding tissue from separate reservoirs in a device the size of a grain of rice.
Those tissues, along with the device itself, were then removed at the end of the procedure, and subsequently inspected for molecular indicators of drug action. So far, the data collected haven’t been used to guide treatments, but retrospective analyses hinted at potential benefits if they had. Two companies — Boston-based Kibur Medical, co-founded by Jonas, and Presage Biosciences, headquartered in Seattle — are now developing these kinds of in situ drug-testing platform.
A choice opportunity
An assay’s treatment predictions are only as good as a patient’s ability to access the recommended drugs — and, when those are expensive cancer agents that have not been approved for the desired use, costs and insurance reimbursement can be impediments.
Pamela Becker, a haematologist at City of Hope cancer centre in Duarte, California, has encountered some of these problems when trying to prescribe drugs that were identified during assay-guided treatment trials for people with multiple myeloma and other blood cancers. “I couldn’t get my top choice,” she says. Becker had to go down the list of recommendations, eventually finding drugs that would be covered by insurance.

The mice with human tumours: Growing pains for a popular cancer model
Another financial obstacle remains reimbursement for the functional tests themselves. In the United States, an official policy enacted in 1996 classifies drug-sensitivity assays as ‘experimental’, making them ineligible for coverage under Medicare, the federal government’s giant health insurance programme for older people. Changing reimbursement rules will thus require reversing that decades-old decision, says Bruce Yeager, an independent consultant in functional precision diagnostics based in Johns Creek, Georgia — an extra hurdle that means “we’re not starting from a point of neutrality”, he says. “We’re starting from negativity.”
Combating such policies and entrenched practices hinges on the availability of compelling clinical data. But accumulating such data can be challenging when the medical establishment is not geared towards enabling functional drug testing. It’s something of a catch-22, says Letai. “But that cycle is going to break in the next couple of years,” he says, “and then I think you’re going to see a sort of non-linear adoption of these strategies, because the power and the need for them is so great.”
Functional testing strategies might even work for conditions outside the cancer arena. In cystic fibrosis, for example, organoid models made from rectal or intestinal tissue are beginning to help clinicians to find effective drug regimens for people with rare disease-causing mutations who are not eligible to receive any approved treatments. “It just makes a lot of sense,” says Jeffrey Beekman, a cystic-fibrosis researcher at the University Medical Center Utrecht in the Netherlands, who has pioneered the approach.
Many cancer researchers feel the same way, and now they just need to prove it to the wider medical community. All eyes are therefore on Staber and his randomized trial, which researchers anticipate will go a long way towards convincing clinicians that genomics is not the be-all and end-all of personalized care. “Paradigm shifts can be very threatening to people,” says Howard, the University of Mississippi radiologist, “but it shouldn’t be threatening. It’s just another tool in our arsenal against disease.”
[ad_2]
Source link