[ad_1]
Life as a scientist in Latin America isn’t always easy — and this is especially true for women. Latina researchers have had to find creative ways to bypass the gender gap from an early age: at 15, girls are half as likely as boys to expect that they’ll work in a science, technology, engineering and mathematics (STEM) area. According to the United Nations, in 2016, less than half (45%) of Latin America’s workforce in research and development were women. Although this figure is above the world average (38%), it is low compared with graduation rates for women in Latin American countries.
Nature spoke to four Latin American researchers about the peaks and troughs they have faced in their careers and how they are connecting science and policy.
ILIANA CURIEL: A translator between cultures
Paediatrician and researcher at the Colombian Institute of Family Welfare in La Guajira, Colombia.
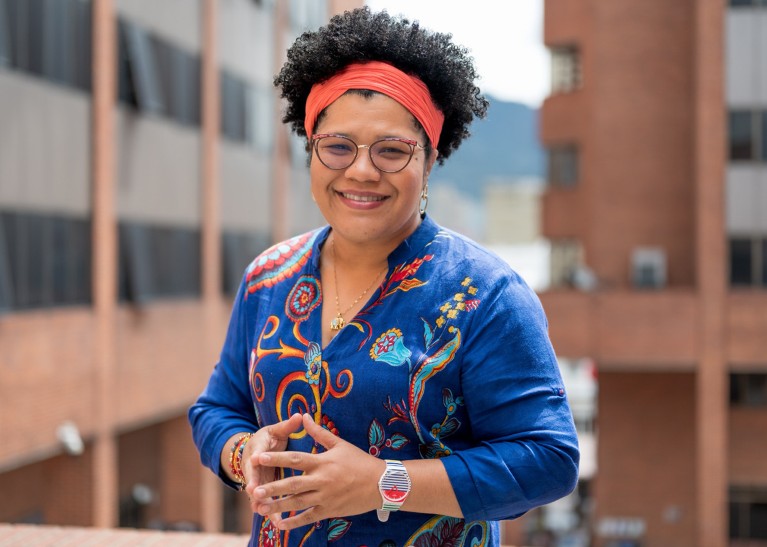

Iliana Curiel became a paediatrician to help her local community in La Guajira, Colombia.Credit: Iliana Curiel Arismendy
I am a mix of Black and Wayuu. I grew up in Uribia, a municipality located in La Guajira, Colombia, at the northernmost tip of South America.
Health challenges in La Guajira are different from those in the rest of Colombia. Although non-transmittable diseases such as obesity and heart conditions demand a considerable effort from public-health services in large cities, the greatest issue in La Guajira is child malnutrition. A child in the region is 60 times more likely to die from undernourishment than is a child living in Bogotá. Most of La Guajira is a desert and access to water can be limited. In some parts of the region, water services reach less than 10% of the population. Around 40% of the population in the territory is under 19, so there is an immense need for paediatric care there. Like other rural and Indigenous communities in Colombia and South America, this is a place where the ‘multidimensional poverty index’ is high and preventable infant and mother mortality abounds.
Growing up in La Guajira, I decided to be a paediatrician and help my community. But I also wanted more than that: after my medical degree in the mid-1990s, I went on to study public health and social policy.

Nature Spotlight: Women in Latin America
Indigenous communities in La Guajira do not easily accept Western medicine because their cultural practices differ from those in urban settings — and public-health policies rarely meet on common ground with these cultural singularities. So, in 2018, I went back there and, together with my wife, started a non-governmental organization, Los Hijos del Sol (Children of the Sun). Our goal has been to conduct research by listening to Indigenous communities, allowing us to plan more adequate models of health care.
Once, for example, we needed to care for a severely undernourished boy. But to offer proper care, we needed to take him from the community to a hospital — and to do that, we had to ask for permission from the community leaders. Because we were in a matrilineal-led group — in which the line of descent is considered from the mothers’ side — it was the boy’s maternal uncles, not his parents, who spoke for the child. So we had to contact his uncle first. A health team, unaware of this, might have asked the boy’s mother for authorization and had a hard time gaining it. If the family think a certain disease is rooted in a spell or bad spirits, we can’t say it’s nonsense — we must adapt our approach and find a shared understanding.
At Los Hijos del Sol, we train Indigenous mothers and midwives to take steps to reduce child mortality. We ask mothers how they know when their child is in trouble, and they come up with the most beautiful analogies. They won’t say the child is “breathing quickly”, but that the child is in a “high tide”, as if the chest and abdomen were moving like restless waves — and they know that it is a sign of alarm.
Most physicians avoid politics, but public health is a political matter and we must be aware of that if we ever want to change things for the better. I’d tell young Latina researchers to never lose sight of your purpose. The path in science, to women, is one of perseverance and resistance, but also of transformation. The qualities that are said to disqualify us as scientists — such as empathy and creativity — are the ones we should take most pride in.
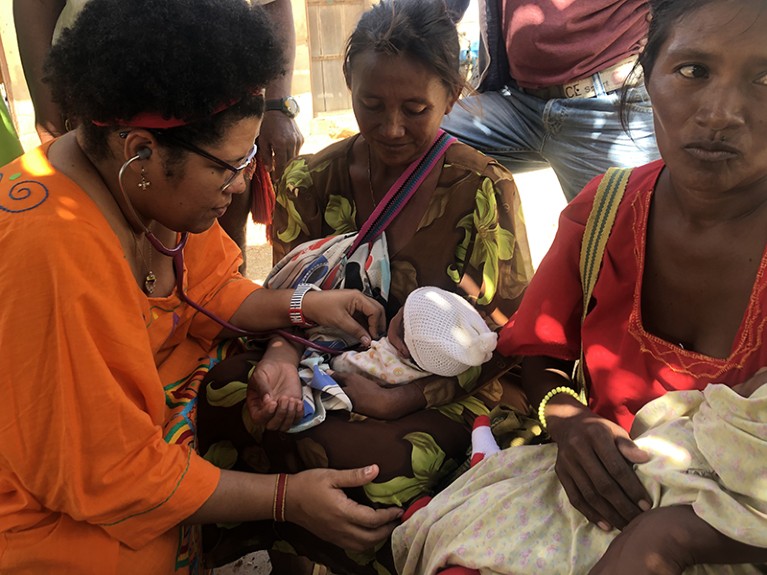

Child malnutrition in La Guajira is one of the biggest issues in the region, says Iliana Curiel (left).Credit: Organización Los Hijos del Sol
XÓCHITL CASTAÑEDA: A voice to Latin American immigrants
Programme director and professor in the School of Public Health at the University of California, Berkeley.
Around 30 years ago, I moved from Mexico City to the United States for my postdoctoral research and it was here that I first saw the negative health impacts felt by immigrants. In the early 2000s, a large number of the migrant community came from Mexico and Latin America. Although the number of migrants from other countries has grown, Mexicans are still the main immigrant workforce in the United States — we’re about 10.6 million people.
During my research at the University of California in San Francisco, I visited the fields where farm workers were employed, and it completely changed my life. I saw the terrible conditions in which they were living to perform the most dangerous, belittling and dirty jobs.

How Latin American researchers suffer in science
I am a medical anthropologist; in the mid-1990s, I was conducting research on the risks that immigrants faced regarding HIV and AIDS. After witnessing neglect and abuse of migrant workers, I realized I couldn’t just stay in academia — I needed to translate research into public action. And this was the beginning of the Health Initiative of the Americas, a programme on health and migration at the University of California, Berkeley.
Since its inception in 2001, the programme has relied on around 20,000 volunteers working to grow a grass-roots movement. I was very fortunate to be part of the University of California system: it helped me to knock at the door of the Mexican government. Because of the magnitude of the Mexican diaspora in the United States, the Mexican government has 50 consulates in the United States. The Mexican government partnered with the programme, and this has opened the doors to cooperation with other Latin American countries, such as Guatemala, El Salvador and Honduras. In the United States, health is unfortunately not a human right — it is sometimes seen as a commodity. We want to extend access to health care to immigrants, who are excluded from the health system, to help improve their living conditions.
We wanted to hold National Health Weeks, just like the ones in Mexico — when the government mobilizes health personnel across the country to knock at houses to give everyone a chance to get vaccinated three times a year. But without accredited health providers, that wouldn’t be possible in the United States. So, we sought out community clinics, and many other organizations started to join: our network has several partners nationwide, including health and cultural institutions and consulates. These are places where immigrants, regardless of their legal status, can access basic health services and advice. Even in remote regions of the United States, they can get vaccines and education about preventive health to improve their overall quality of life.
Young Latina researchers have the opportunity and the responsibility to contribute to a more equitable world. My advice is to never give up. Even in hard times there is light, and public health is a marvellous instrument to shine that light.
DENISE LAPA: A fetoscopy pioneer
Fetal and neonatal surgery programme coordinator at Sabará Child Hospital in São Paulo, Brazil.
In 1999, I started to develop a technique to treat spina bifida — a pre-birth condition in which the neural tube bulges on the back of the fetus. The condition can damage nerves in the spinal cord and greatly affect a child’s ability to walk or perform day-to-day activities.
In the late 1990s, Thomas Kohl, who is now head of the German Center for Fetal Surgery and Minimally-Invasive Therapy at the University Medical Center Mannheim, developed a technique to close the gap that forms in the spine. His idea was to stitch the fetus’s spine without opening the mother’s womb. I had been testing a similar technique for a decade when, in 2012, he invited me to Germany. We started an informal exchange.
The difference between Kohl’s technique and mine was that, instead of stitching all of the layers in the back of a fetus — spinal cord, muscle and skin tissues — my team and I used a biocellulose patch over the spinal tissue to help it self-heal and avoid suturing the fetus’s spinal cord to the tissue above it.

Affirmative action slow to take hold in Brazil’s graduate science education
Throughout my career, I felt I had to prove myself all the time as a woman and, as a Latina researcher, I also had regional prejudice on top of that. To me, it seemed that some people, most of whom were men, felt that if a breakthrough in fetoscopy (fetal endoscopy) was to be made, it wouldn’t be made by a woman and certainly not one from Brazil.
However, in 2013, after 14 years of testing in animal models, our first fetal surgery at the Samaritan Hospital of São Paulo proved that the technique worked. A decade later, we could see that not only was it viable, but also that it yielded positive long-term results. A study1 following 78 children who had undergone our procedure showed that almost half of them (46%) could walk independently once they reached between 2.5 and 10 years old — and almost all of them (94%) had expected social function. In comparison, a 2020 study2 on the effectiveness of the conventional open-womb surgical technique showed that around 29% of children aged 6 and over who had undergone this surgery could walk independently. Previous studies have shown that the effectiveness of the conventional technique in terms of walking rates is as high as 45%3.
As well as in Brazil, our technique is now used in Israel, Chile, Uruguay, Italy and parts of the United States. It’s also rising in popularity: more than 300 surgeries have been performed outside Brazil. Everything I did in my life, I accomplished because a man told me I couldn’t. It’s extremely rewarding to see children, whose parents relied on my team, being able not only to walk, but also to jump and play freely — some even go skiing and do ballet.
My piece of advice to young Latina researchers would be: structural sexism is still not understood by most men. It is up to us, women, to occupy important spaces and teach our daughters a different language of love and respect between men and women.
YESTER BASMADJIÁN:On the front line against insect-borne diseases
Head of the Department of Parasitology and Mycology in the Medicine Faculty at the University of the Republic in Montevideo, Uruguay.
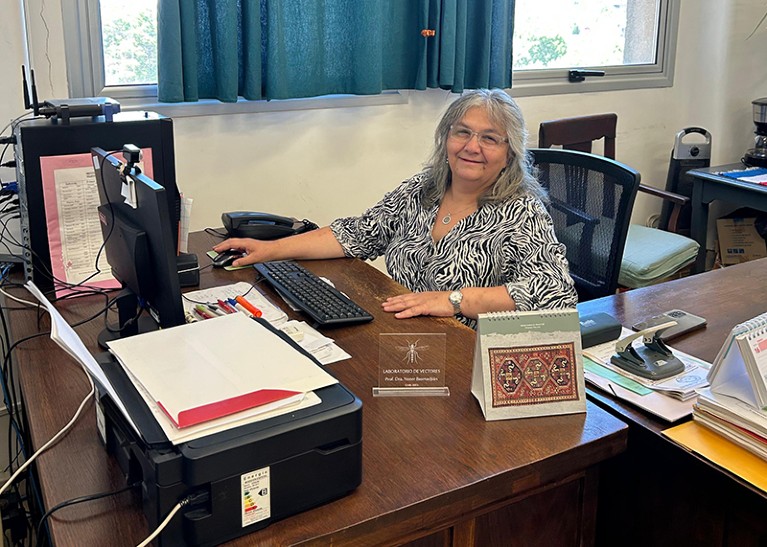

Yester Basmadjián says protecting against misinformation is an important part of her job.Credit: Ramiro Tomasina
Before the viral disease dengue returned to Uruguay in 2016, the last epidemic had been a century earlier, in 1916. In the late 1950s, the country had eradicated the mosquito vector Aedes aegypti through monitoring populations and their behaviour. But, because the continent never fully got rid of it, the mosquito returned in 1997. Despite heavy public campaigning, the country was unable to eradicate it again. Now we’re seeing a rise in local transmission of dengue, especially in the Montevideo region and Salto on the border with Argentina. There were 48 confirmed cases in 2023, and this year we have seen more than 700.
Cases of dengue, most of which were imported by travellers from neighbouring countries such as Brazil, Argentina and Paraguay, are now a concern in Montevideo. At the University of the Republic in Montevideo, we have a laboratory in which we can study this and other disease-vector insects more closely. Our lab has the support of Uruguay’s Public Health Ministry, the International Atomic Energy Agency (IAEA) and the Pan American Health Organization, and we have partnered with a number of institutions in Brazil and other Latin American countries.
We’re using X-rays (hence our partnership with the IAEA) to sterilize male A. aegypti mosquitoes before they become adults, to decrease their overall population. Female mosquitoes mate only once; if they mate with sterile males then they won’t produce offspring. Another advantage is that male mosquitoes generally don’t interact with people and, because they do not feed on blood, they don’t transmit diseases. Our project will not eliminate this insect in Uruguay, but it’s a tool that will add to the fight. It is certainly better than open-air insecticide spraying — we don’t know whether mosquitoes in Uruguay are resistant to certain chemicals. We’re launching close to 30,000 first-generation sterile mosquitoes at the end of this year and are looking forward to good results.
One of our biggest challenges is ensuring that the new lab remains operational in both the medium and long term — not only by maintaining resources, but also by protecting against a wave of misinformation and conspiracy theories. Many people think that sterilization of mosquitoes is going to cause a change in human bodies (which is not possible even if a male mosquito interacted with a person). At the lab, we try to counter this through outreach with journalists and by promoting workshops in schools.
Although sterilizing mosquitoes is not a silver bullet to end dengue, it’s an important tool, and the public’s cooperation is essential to fight the mosquito that transmits it.
My advice to young Latina researchers is that we have to study a lot to adapt to an ever more technological world — but it’s important never to give up when faced with challenges. Always move forward and, at some point, you’ll get to where you want to be.
[ad_2]
Source link