[ad_1]

Researchers have leveraged artificial intelligence to enhance the photostability of molecules for solar energy applications, achieving molecules four times more stable than previous ones.
Their novel approach involved AI-driven closed-loop experimentation and automated chemical synthesis to uncover the underlying chemical principles of stability, offering fresh insights into molecular design for organic solar cells.
Artificial intelligence is a powerful tool for researchers, but with a significant limitation: The inability to explain how it came to its decisions, a problem known as the “AI black box.” By combining AI with automated chemical synthesis and experimental validation, an interdisciplinary team of researchers at the University of Illinois Urbana-Champaign has opened up the black box to find the chemical principles that AI relied on to improve molecules for harvesting solar energy.
Advancements in Light-Harvesting Molecule Stability
The result produced light-harvesting molecules four times more stable than the starting point, as well as crucial new insights into what makes them stable — a chemical question that has stymied materials development.
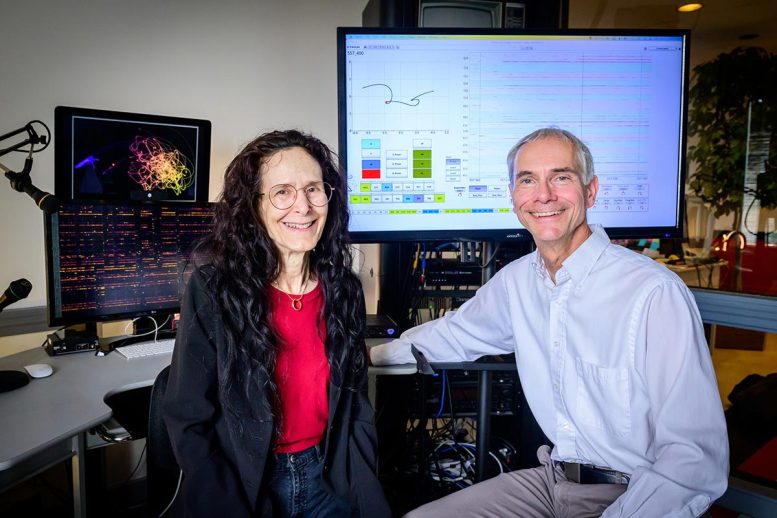
The interdisciplinary team of researchers was co-led by U. of I. chemistry professor Martin Burke, chemical and biomolecular engineering professor Ying Diao, chemistry professor Nicholas Jackson and materials science and engineering professor Charles Schroeder, in collaboration with along with University of Toronto chemistry professor Alán Aspuru-Guzik. They published their results today (August 28) in the journal Nature.
“New AI tools have incredible power. But if you try to open the hood and understand what they’re doing, you’re usually left with nothing of use,” Jackson said. “For chemistry, this can be very frustrating. AI can help us optimize a molecule, but it can’t tell us why that’s the optimum — what are the important properties, structures and functions? Through our process, we identified what gives these molecules greater photostability. We turned the AI black box into a transparent glass globe.”

Solving Photostability With Closed-Loop Experimentation
The researchers were motivated by the question of how to improve organic solar cells, which are based on thin, flexible materials, as opposed to the rigid, heavy, silicon-based panels that now dot rooftops and fields.
“What has been hindering commercialization of organic photovoltaics is problems with stability. High-performance materials degrade when exposed to light, which is not what you want in a solar cell,” said Diao. “They can be made and installed in ways not possible with silicon and can convert heat and infrared light to energy as well, but the stability has been a problem since the 1980s.”
Accelerating Discovery with Modular Chemistry and AI
The Illinois method, called “closed-loop transfer,” begins with an AI-guided optimization protocol called closed-loop experimentation. The researchers asked the AI to optimize the photostability of light-harvesting molecules, Schroeder said. The AI algorithm provided suggestions about what kinds of chemicals to synthesize and explore in multiple rounds of closed-loop synthesis and experimental characterization. After each round, the new data were incorporated back into the model, which then provided improved suggestions, with each round moving closer to the desired outcome.
The researchers produced 30 new chemical candidates over five rounds of closed-loop experimentation, thanks to building block-like chemistry and automated synthesis pioneered by Burke’s group. The work was done at the Molecule Maker Lab housed in the Beckman Institute for Advanced Science and Technology at the U. of I.
“The modular chemistry approach beautifully complements the closed-loop experiment. The AI algorithm requests new data with maximized learning potential, and the automated molecule synthesis platform can generate the new required compounds very quickly. Those compounds are then tested, the data goes back into the model, and the model gets smarter — again and again,” said Burke, who also is a professor in the Carle Illinois College of Medicine. “Until now, we’ve been largely focused on structure. Our automated modular synthesis now has graduated to the realm of exploring function.”
Unveiling the Secrets of Molecular Stability
Instead of simply ending the query with the final products singled out by the AI, as in a typical AI-led campaign, the closed-loop transfer process further sought to uncover the hidden rules that made the new molecules more stable.
As the closed-loop experiment ran, another set of algorithms was continuously looking at the molecules made, developing models of chemical features predictive of stability in light, Jackson said. Once the experiment concluded, the models provided new lab-testable hypotheses.
“We’re using AI to generate hypotheses that we can validate to then spark new human-driven campaigns of discovery,” Jackson said. “Now that we have some physical descriptors of what makes molecules photostable, that makes the screening process for new chemical candidates dramatically simpler than blindly searching around chemical space.”
To test their hypothesis about photostability, the researchers investigated three structurally different light-harvesting molecules with the chemical property they identified — a particular high-energy region — and confirmed that choosing the proper solvents made the molecules up to four times more light-stable.
“This is a proof of principle for what can be done. We’re confident we can address other material systems, and the possibilities are only limited by our imagination. Eventually, we envision an interface where researchers can input a chemical function they want and the AI will generate hypotheses to test,” Schroeder said. “This work could only happen with a multidisciplinary team, and the people, resources, and facilities we have at Illinois, and our collaborator in Toronto. Five groups came together to generate new scientific insight that would not have been possible with any one of the sub-teams working in isolation.”
Reference: “Closed-loop transfer enables AI to yield chemical knowledge” 28 August 2024, Nature.
DOI: 10.1038/s41586-024-07892-1
This work was supported by the Molecule Maker Lab Institute, an AI Research Institutes program supported by the U.S. National Science Foundation under grant no. 2019897.
[ad_2]
Source link