Experimental procedures
Tissue samples
Heart samples were collected in strict observance of the legal and institutional ethical regulations. The heart samples were collected under a University of California San Diego (UCSD) Human Research Protections Program Committee Institutional Review Board (IRB)-approved protocol (IRB number 081510) by the UCSD Perinatal Biorepository’s Developmental Biology Resource after informed consent was obtained from the donor families. All experiments were performed within the guidelines and regulations set forth by the IRB (IRB number 101021, registered with the Developmental Biology Resource). Ethical requirements for data privacy include that sequence-level data (for example, fastq files) be shared through controlled-access databases.
Tissue processing
Tissue samples were collected in buffer containing 10 mM HEPES pH 7.8, 130 mM NaCl, 5 mM KCl, 10 mM glucose, 10 mM BDM, 10 mM taurine, 1 mM EDTA and 0.5 mM NaH2PO4, and overall morphology was checked under a stereotaxic dissection microscope (Leica).
For single-cell dissociation, tissue samples from eight hearts were further cut into small pieces and enzymatically digested by incubating with collagenase type IV (Gibco) and Accutase (ThermoFisher) at 37 °C for 60 min. After removing the dissociation medium, cells were resuspended in PBS supplemented with 5% FBS and sorted using a Sony SH800 sorter. Samples were diluted to approximately 1,000 cells per µl before processing for scRNA-seq, as shown in Supplementary Fig. 1a.
Samples for MERFISH were washed with ice-cold PBS and then fixed in 4% paraformaldehyde at 4 °C overnight. On the second day, samples were washed in ice-cold PBS 3 times, 10 min each, and were incubated in 10% and 20% sucrose at 4 °C for 4 h each, and in 30% sucrose overnight, followed by immersion with OCT (Fisher, 23-730-571) and 30% sucrose (v/v) for 1 h. The samples were then embedded in OCT and stored at −80 °C until sectioning.
hPSCs
For the single-cell and bioprinting studies, a H9-hTnnTZ-pGZ-D2 hPSC line (TNNT2:eGFP hPSC cardiomyocyte reporter line) was purchased from WiCell and maintained as previously described41. For the bioprinting studies, an additional engineered TNNT2:NLS-mKATE2 RUES2 hPSC cardiomyocyte transgenic reporter line that specifically expresses the mKATE2 fluorescent protein containing a nuclear localization signal (NLS-mKATE2) in differentiated cardiomyocytes was used (Supplementary Fig. 18a). Both lines were routined authenticated with fluorescence microscopy, immunofluorescence and flow cytometry studies, and tested negative for mycoplasma contamination by PCR. To generate the TNNT2:NLS-mKATE2-T2A-BsdR RUES2 hPSC cardiomyocyte reporter line (TNNT2:NLS-mKATE2), we transfected a RUES2 hPSC line with a Piggybac (PB) construct expressing NLS-mKATE2-T2A-BsdR driven by the cardiomyocyte-specific TNNT2 promoter. To clone the PB-TNNT2:NLS-mKATE2-T2A-BsdR, we used the PB plasmid pcsj532 (a gift from K. Willert, UCSD) and used Gibson assembly (SGI, GA1200) to clone in a synthesized TNNT2 promoter42 (Integrated DNA Technologies), PCR-amplified NLS-mKATE2-T2A-BsdR (with polyA) from pgRNA-CKB43 (a gift from B. Conklin, Gladstone; Addgene, plasmid 73501) and PCR-amplified PGK:PuroR from RT3GEPIR44 (a gift from J. Zuber, IMP, Austria; Addgene, plasmid 111169). All three components were assembled in one Gibson assembly with pcsj532 digested using NheI (NEB R3131L). RUES2 hPSCs were transfected using Lipofectamine STEM (Invitrogen, STEM00015) with the PB-TNNT2:NLS-mKATE2-T2A-BsdR and a plasmid expressing a human-optimized PB transposase (pcsj533, a gift from K. Willert, UCSD) to integrate the PB. Two days after transfection, the cells were selected using 0.4 μg ml–1 puromycin. The subsequent surviving cells behaved similarly to the parental and the TNNT2:eGFP hPSC lines in terms of proliferation and differentiation. Protocols were approved by the IRB (number 190561) at UCSD.
hPSC cardiac cell differentiations and sample preparation
hPSC lines were cultured in E8 medium and grown on Geltrex (Gibco)-coated plates. Differentiation of hPSCs into cardiomyocytes was performed using established protocols as previously described41,45,46. In brief, hPSCs were grown to 80% confluency, and on day 0 (D0), cells were cultured with RPMI/B27 supplement without insulin (B27 minus insulin; ThermoFisher) containing 10 µM CHIR (Fisher Scientific). After 24 h of CHIR application, the medium was replaced with fresh B27 without insulin and the cells were cultured for another 48 h. Next (D3), 5 µM IWP2 (Tocris) was supplemented to B27 without insulin and cultured for another 48 h. At D5, the B27 without insulin and with IWP2 was replaced with fresh B27 without insulin for another 48 h. From D7 onwards, cells were maintained in RPMI/B27 with insulin (B27, ThermoFisher). On D15, this B27 medium was then supplemented with either NRG1 (50 ng ml–1)47 or PBS, and further cultured until D30 and greater, refreshing the medium every 3 days.
scRNA-seq studies performed on hPSC-derived samples were prepared as described in the ‘Tissue processing’ section. In brief, D25 hPSC-derived cardiac cells were enzymatically digested by incubating with collagenase type IV (Gibco) and Accutase (ThermoFisher) at 37 °C for 60 min. After removing the dissociation medium, cells were resuspended in PBS supplemented with 5% FBS and sorted using a Sony SH800 sorter.
Animal studies
Animal studies were conducted in strict compliance with the Guide for the Care and Use of Laboratory Animals published by the National Institutes of Health and protocols approved by the Institutional Animal Care and Use Committee of UCSD (A3033-01). Mice were maintained on a 12 h–12 h light–dark cycle in a controlled temperature (20–22 °C) and humidity (30–70%) environment. The generation of Tcf21-creERT2 and Sema3cfl/fl mice has been previously described48,49. To validate the genotype of the mice, genomic DNA was extracted by adding a 2 mm tail clipping to a 75 µl solution containing 25 mM NaOH and 0.2 mM EDTA, and then heating the sample for 30 min at 98 °C. Next, 75 µl of 40 mM Tris-HCl (pH 5.5) was then added to neutralize the reaction, and a 1:50 dilution of genomic DNA template was used for genotyping PCR. Both male and female embryos were used in this study; the embryos were not genotyped to determine sex. To determine the developmental stage of embryonic development during which tamoxifen treatment was administered, noon on the day of the vaginal plug was assumed to be E0.5. Tamoxifen (Sigma, T5648-1G, 0.1 mg g–1 body weight) was fed to pregnant mice by gavage at E10.5, and hearts were collected at E12.5, E14.5, E17.5 and postnatal day 1. The fixed hearts were embedded in paraffin, sectioned and stained with haematoxylin and eosin by the UCSD Histology Core. Images were taken on a Hamamatsu Nanozoomer Slide Scanning system and an Olympus VS200 slide scanner, and processed using NDP View 2 software (Hamamatsu) and QuPath (v.0.4.3)50, respectively. Phenotypic analyses of ventricular wall thickness were performed as previously described51. In brief, the thicknesses of ventricular compact and trabecular zones were measured at the level of the papillary muscle, with measurements taken from at least three areas per section, and at least three sections per mouse.
Single-cell transcriptome library preparation and sequencing
Single-cell droplet libraries using the cell suspensions from the Sony SH800 sorter were prepared according to the manufacturer’s instructions using the 10x Genomics Chromium controller, Chromium Single Cell 3′ Library and Gel Bead kit v2 (PN-120237) and Chromium i7 Multiplex kit (PN-120262). All libraries were sequenced on a HiSeq 4000 (Illumina) to a mean read depth of at least 65,000 total aligned reads per cell.
MERFISH gene selection
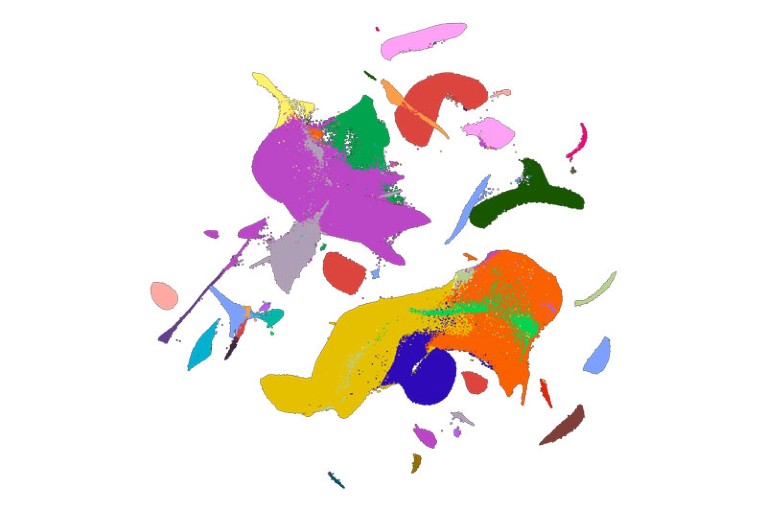
To spatially detect cell populations identified in the scRNA-seq dataset, we designed a panel of 238 genes specific for these subpopulations. These genes were then simultaneously imaged on cardiac samples using the combinatorial barcoded imaging technique MERFISH7. We initially identified gene markers differentially expressed for each of the 75 cell subpopulations by performing differential gene expression (DGE) analyses as well as applying a NS-Forest2 (ref. 52) classifier on scRNA-seq data obtained from the aforementioned human hearts in Supplementary Fig. 1. All markers were combined from the binary gene analysis utilizing NS-Forest2 (ref. 52) (159 genes) (Supplementary Table 7) and DGE analysis (7,557 genes) (Supplementary Table 3) of the cell subpopulations, and were then filtered for genes that were either not long enough to construct 48 target regions (each 30-nucleotides long) without overlap or for which expression levels were outside the range of 0.01–300 average unique molecular identifier (UMI) per cluster, as measured by scRNA-seq. The performance of identifying marker genes between NS-Forest2 and Spapros53 pipelines was also compared. The initial result of Spapros produced 80 genes, which is half the number chosen by NS-Forest2. To compare a similar number of genes between NS-Forest2 and Spapros, these 80 genes were removed from the dataset and Spapros was run again, which selected another 90 genes. The combination of these two sets of genes were used for the Spapros gene list (Supplementary Table 9). To quantify the ability of the selected genes to re-identify cell subpopulations at the same granularity as annotated in the scRNA-seq data, the dimensionality reduction and neighbour graph were recalculated using only the genes selected by the algorithm (NS-Forest2 or Spapros) that was being evaluated. Each cell was then reassigned its cell subpopulation label based on the majority cell subpopulation of its five nearest neighbours in the new neighbour graph. The percentage of cells reassigned their original label was used as an accuracy metric. With this metric, we found that NS-Forest2 and Spapros chose genes with similar performance (Supplementary Table 10). Among the 238 MERFISH target genes, 63 were manually selected from the DGE and NS-Forest2 gene lists, including established markers for atrial, ventricular and non-chambered cardiomyocytes, as well as non-cardiomyocyte cell markers for fibroblasts, pericytes, VSMC, epicardial, endocardial, BEC, LEC and immune cells. Genes specific for platelet–red blood cells were not selected. To validate the final target gene list, we tested whether we could transcriptionally rederive the cell populations by cluster analyses using only the 238 target genes. To this end, we reduced the scRNA-seq dataset to only the 238 genes in the MERFISH gene panel and then performed dimensionality reduction, graph-based clustering and UMAP visualization. We observed a similar level of transcriptional separation and definition of cell classes between using the 238 target genes versus using the 3,000 variable genes chosen to annotate the cell classes in the scRNA-seq data (Fig. 1a and Extended Data Fig. 1b). In addition to the 238 MERFISH genes, we selected 11 genes that were imaged sequentially using smFISH (Supplementary Table 11), including genes that validated the combinatorial MERFISH imaging (Extended Data Fig. 2d).
MERFISH probe library design and construction
A total of 238 genes were identified as MERFISH target genes, which were subsequently used for probe generation and MERFISH assays as shown in Supplementary Table 11. To encode MERFISH RNA probes for spatial detection, a 22-bit modified Hamming distance 4 code was used8. Each of the 238 possible barcodes required at least 4 accumulated errors to be converted into another barcode. This property permitted the detection of errors up to any 2 bits, and the correction of errors to any single bit. In addition, this encoding scheme used a constant Hamming weight (that is, the number of 1 bits in each barcode) of 4 to avoid potential bias in the measurement of different barcodes due to a differential rate of 1 to 0 and 0 to 1 errors and because the optical density within each bit can interfere with resolving individual fluorescent spots, as previously described6. We used 238 out of the 7,315 possible barcodes to encode cellular RNAs and chose 10 barcodes that were left unassigned to serve as blank controls. The encoding probe set that we used contained 30–48 encoding probes per RNA, with each encoding probe containing 3 out of the 4 readout sequences assigned to each RNA. Encoding probes were designed using our own pipeline, namely, ProbeDesign. ProbeDesign was developed using a fully optimized algorithm in C++ for both DNA and RNA probes. ProbeDesign used the same principles of probe design utilized by various published algorithms (OligoArray54, OligoMiner55, OligoPaint56 and ProbeDealer57), for which off-targets are based on genome-wide 17-nucleotide off-target counts. Probes were selected with similar GC content or melting temperature, and the repetitive regions were used for off-target counting but not for probe design. ProbeDesign was implemented in three steps. (1) Build a 17-nucleotide index based on the reference genome (DNA) or genome annotation files (RNA). This step is fully optimized with bit-vector and hash tables for high-performance computation; (2) Scan selected loci or genome sequences to calculate the off-targets based on the 17-nucleotide counts in the previous step. And (3), filter and rank probe candidates based on predefined selection criteria. For the RNA probe design, we used the transcript sequences derived from the human reference genome sequences (hg38) downloaded from ncbi_refseq (https://hgdownload.soe.ucsc.edu/goldenPath/hg38/bigZips/genes/). The generation of the encoding probe sets were prepared from oligonucleotide pools, as previously described58,59. In brief, we first used limited-cycle PCR to amplify the oligopools (Twist Biosciences). Then, we used these DNA sequences as the templates for in vitro transcription into RNA using T7 polymerase (NEB, E2040S). Subsequently, the RNA products were converted into single-stranded DNA with Maxima Reverse Transcriptase enzyme (Thermo Scientific, EP0751), and then the DNA was purified by alkaline hydrolysis (to remove the RNA templates) followed by DNA oligo purification kits (Zymo Research, D4060).
MERFISH sample preparation
Frozen hearts were sectioned at −20 °C on a cryostat (Leica CM3050S). A series of 12 μm coronal slices were cut at about 600 μm along the anterior–posterior axis of collected human hearts, which captured all of the major cardiac structures. MERFISH measurements of 238 genes with 10 non-targeting blank controls were performed as previously described using the encoded barcode sequences (Supplementary Table 11) and published readout probes60. In brief, 12-μm-thick tissue sections were mounted on 40 mm no. 1.5 coverslips that were silanized and poly-l-lysine coated58 and subsequently pre-cleared by immersing into 50% (v/v) ethanol, 70% (v/v) ethanol and 100% ethanol, each for 5 min. The tissue was then air-dried for 5 min, followed by treatment with Protease III (ACDBio) at 40 °C for 30 min, and then washed with PBS for 5 min. Tissues were then preincubated with hybridization wash buffer (30% (v/v) formamide in 2× SSC) for 10 min at room temperature. After preincubation, the coverslip was moved to a fresh 60 mm Petri dish, and the residual hybridization wash buffer was removed with a Kimwipe laboratory tissue. In the new dish, the coverslip was immersed with 50 μl of encoding probe hybridization buffer (2× SSC, 30% (v/v) formamide, 10% (w/v) dextran sulfate, 1 mg ml–1 yeast tRNA and a total concentration of 5 μM encoding probes and 1 μM of anchor probe: a 15-nucleotide sequence of alternating dT and thymidine-locked nucleic acid (dT+) with a 5′-acrydite modification (Integrated DNA Technologies)). The sample was then placed in a humidified 37 °C oven for 36–48 h and then washed with hybridization wash buffer for 20 min at 37 °C and 20 min at room temperature. Samples were post-fixed with 4% (v/v) paraformaldehyde in 2× SSC and then washed with 2× SSC with murine RNase inhibitor for 5 min. To anchor the RNAs in place, the encoding probe-hybridized samples were embedded in thin, 4% polyacrylamide (PA) gels as previously described58. In brief, the hybridized samples on coverslips were first washed with a de-gassed 4% PA solution, consisting of 4% (v/v) of 19:1 acrylamide/bis-acrylamide (Bio-Rad, 1610144), 60 mM Tris⋅HCl pH 8 (ThermoFisher, AM9856), 0.3 M NaCl (ThermoFisher, AM9759) and a 1:1,000 dilution of 0.1-µm-diameter blue fluorescent (350/440) beads (Life Technologies, F-8797). The beads served as fiducial markers for the alignment of images taken across multiple rounds of imaging. The coverslips were then washed again for 2 min with the same 4% PA gel solution supplemented with the polymerizing agents ammonium persulfate (Sigma, A3678) and TEMED (Sigma, T9281) at final concentrations of 0.03% (w/v) and 0.15% (v/v), respectively. The gel was then allowed to cast for 1.5 h at room temperature. The coverslip and the glass plate were then gently separated, and the PA film was incubated with a digestion buffer consisting of 50 mM Tris⋅HCl pH 8, 1 mM EDTA, 0.5% (v/v) Triton X-100 in nuclease-free water and 1% (v/v) proteinase K (New England Biolabs, P8107S). The sample was digested in this buffer for >36 h in a humidified, 37 °C incubator and then washed with 2× SSC 3 times. The samples were finally stained with an Alexa 488-conjugated anchor probe-readout oligonucleotide (Integrated DNA Technologies) and DAPI solution at 1 μg ml–1. MERFISH measurements were conducted on a home-built system as previously described60.
Immunofluorescence studies
For the immunofluorescence studies of the TNNT2:NLS-mKATE2 hPSC line, D25 hPSC-derived cardiac cells were dissociated, replated and then cultured for another 4 days before being fixed. The fixed cells were then immunostained, as previously described41, using an antibody for TNNT2 (A647 mouse anti-cardiac troponin T, BD 565744, 1:200). Nuclei were visualized using DAPI (1 µg ml–1, Roche) staining. Immunofluorescent images were taken on a Nikon C2 confocal microscope.
Real-time quantitative PCR
RNA expression was measured by quantitative PCR (qPCR). RNA was extracted using TRIzol reagent (ThermoFisher) and a Direct-zol RNA MiniPrep kit (Zymo Research). cDNA was generated using 1,000 ng RNA mixed with iScript Reverse Transcription Supermix (Bio-Rad) and then diluted 1:10 in UltraPure DNase/RNase-free distilled water (ThermoFisher). qPCR was performed using Power SYBR Green master mix (ThermoFisher) according to the manufacturer’s recommendations, and a two-step amplification CFX_2stepAmp protocol on a Bio-Rad CFX Connect Real-Time PCR Detection system. Data were analysed using the 2−ΔΔCt method. All gene expression was normalized to the expression of TATA box binding protein (TBP). Primer sequences are listed in Supplementary Table 22.
In vitro hPSC ventricular wall model
To create an in vitro hPSC ventricular wall model for studying ventricular wall morphogenesis, we bioprinted cardiomyocytes in multilayered constructs as shown in Fig. 5. To this end, we differentiated TNNT2:eGFP and TNNT2:NLS-mKATE2 hPSCs into D15 cardiomyocytes (hPSC-CMs) as described in the ‘hPSC cardiac differentiations and sample preparation’ section. D15 TNNT2:eGFP hPSC-CMs were further treated with NRG1 to create trabecular-like CMs as previously described46. As controls, D15 TNNT2:eGFP and TNNT2:NLS-mKATE2 hPSC-CMs were treated with PBS. D30+ hPSC-CMs (>90% efficiency by flow cytometry) were then enzymatically dissociated with collagenase type IV (Gibco) and Accutase (ThermoFisher) and resuspended at 100 million cardiomyocytes per ml. The method to bioprint multilayered constructs involved printing D30+ hPSC-CMs that were treated with NRG1 or PBS into a rectangle with finger-like projections that was 500 × 700 × 600 µm (height × width × length) (inner-LV CC-like layer), followed by printing an adjacent rectangular structure (500 × 700 × 75 µm) (intermediate-LV CC-like layer) containing gelatin methacryloyl (GelMA)61 mixed with 100 ng ml–1 of different combinations of SEMA3C, SEMA3D, SEMA6A or SEMA6B (R&D Systems) proteins as described in Fig. 5b. The concentration of SEMA3C used for the conditions in Fig. 5c was 1,000 ng ml–1 because SEMA3C was located further from the inner-LV CC-like layer. The cell-encapsulated layer was fabricated using 6.25% GelMA and 0.33% LAP with 15 s of light exposure time, and the cells were mixed with the monomer solution directly before bioprinting. The adjacent layer containing the signalling factors was printed using 4% GelMA and 0.4% LAP with 15 s of light exposure time. Using a methacrylated coverslip fixed to the controller stage, a 20 µl cell–material mixture was placed into the space between the coverslip and a polydimethylsiloxane (PDMS) film attached to a glass slide. This cell–material mixture was then exposed to UV light (365 nm, 88 mW cm–2) with a pattern generated by a digital micromirror device chip. After printing each layer, the construct was washed three times with warm PBS and aspirated dry. Finally, the multilayered construct was washed in both PBS and medium, and then stored in a cell culture incubator (37 °C, 5% CO2). Medium was refreshed every other day.
Data analysis
Processing of raw sequencing reads
Raw sequencing reads were processed using the Cell Ranger (v.3.0.1) pipeline from 10x Genomics. Reads were demultiplexed and aligned to the human hg38 genome, and UMI counts were quantified per gene per cell to generate a gene–barcode matrix.
Cell filtering and clustering
After generating the gene–barcode matrix file from Cell Ranger, the individual count matrices were merged together and processed using the Seurat (v.4.0.1) R package62 (https://satijalab.org/seurat/). Further filtering and clustering analyses of the scRNA-seq cells were performed using the Seurat package, as described in the tutorials (https://satijalab.org/seurat/). Cells with at least 1,000 genes detected and a mitochondrial read percentage of less than 30% were used for downstream processing. Potential doublets were removed using DoubletFinder (v.2.0)63 (https://github.com/chris-mcginnis-ucsf/DoubletFinder) using an anticipated doublet rate of 5%, which is the expected rate reported by 10x Genomics for the number of cells loaded onto the 10x Controller. For the aggregated dataset, gene expression was normalized for genes expressed per cell and total expression using the NormalizeData function. The top 3,000 variable genes were detected using the FindVariableFeatures function with default parameters. All of the genes were subsequently scaled using the ScaleData function, which utilizes a linear regression model to eliminate technical variability due to the number of genes detected, replicate differences and mitochondrial read percentage. Principal components were calculated using RunPCA, and the top 50 principal components (supported by ElbowPlot showing diminishing variance explained beyond the top 50 principal components) were used for creating the nearest neighbour graph utilizing the FindNeighbors function with k.param = 50. The generated nearest neighbour graph was then used for graph-based, semi-unsupervised clustering (FindClusters, default resolution of 0.8) and UMAP to project the cells into two dimensions. Marker genes were identified using a Wilcoxon rank-sum test (FindAllMarkers, default parameters) for one-versus-all comparisons for each of the cell clusters. Cell identities were assigned to the clusters by cross-referencing their marker genes with known cardiac cell type markers from both human and mouse studies, in addition to in situ hybridization data from the literature10,11,12,13,14,15. On occasion, a cell cluster would emerge that expressed marker genes representing multiple populations, as well as contained cells with low UMI and gene counts that escaped the first filtering step. These cells were removed from downstream analyses. The clustering approach was then repeated for each compartment of cells (cardiomyocyte, mesenchymal, endothelial, neuronal and blood) as described above, and the clustering accuracy was evaluated using SCCAF (v.0.0.10)64 with the following parameters: linear regression with L1 regularization with a 50/50 train-test split and a fivefold cross validation. For the adult ventricle scRNA-seq comparison, we combined previously published datasets33 with our developing heart scRNA-seq dataset and re-ran the NormalizeData function to compare gene expression between these datasets.
Label transfer analysis
Cell annotations from the scRNA-seq dataset were compared to a recently published adult heart dataset18 utilizing scArches (v.0.5.9)65. For scArches, both the adult and developing transcriptomic datasets were integrated using scVI (v.1.0.3)66 (n_hidden=128, n_latent=50, n_layers=3, dispersion = ‘gene-batch’). A reference hierarchy tree was then constructed using the treeArches67 workflow (https://docs.scarches.org/en/latest/treeArches_identifying_new_ct.html) with default parameters and “cell_state” labels on the adult heart published reference dataset18. Labels from the reference dataset were then transferred to the developing heart query dataset to predict the cell labels utilizing the scHPL.predict.predict_labels() function with default parameters. Rejected cells were calculated using the posterior probability (default option) with a 0.5 threshold.
Gene regulatory network analysis
To identify age-related changes in gene expression, we applied the pySCENIC (v.0.12.1)68 gene regulatory network (GRN) inference tool to our scRNA-seq dataset. To this end, the scRNA-seq dataset was split by cell class, and pySCENIC analysis was performed to identify cell-class-specific regulons following the standard pipeline on GitHub (https://github.com/aertslab/SCENICprotocol/). In brief, we performed three steps to create a GRN for each cell class: (1) GRN inference using the GRNBoost2 algorithm, (2) transcription factor (TF) regulon predictions and (3) cellular enrichment area under the curve (AUC, measure of regulon activity) calculation for each cell. The resulting AUC matrix was then used to identify the regulons with the most significant change of activity over age by fitting a linear model to regulon activity and identifying regulons with the highest positive and negative rate of change. To find the functional pathways enriched in each set of regulons, we performed gene ontology enrichment analysis using the EnrichR Bioconductor package (v.3.1)69. On the same regulons, we constructed a regulatory network with the top 100 non-redundant edges of the network by importance score, and visualized the edges, transcription factors and target genes using Cytoscape (v.3.8.0)70. For the overall GRN of vCMs, we constructed a regulatory network with the top 50 transcription factors by centrality and then took the top 500 non-redundant edges of the network by importance score and visualized the edges and transcription factors using Cytoscape.
CCI analysis
We applied CellChat (v.1.6.1)71 to our scRNA-seq dataset to identify region-specific CCIs. Atrial cells and ventricular cells were divided based on their region of dissection (LA/RA for atrial and LV/RV/IVS for ventricular) and were analysed separately. Next, we followed the suggested workflow of CellChat, using its database of human ligand–receptor interactions (with the addition of the NRG1–ERBB2 signalling pathway owing to its known biological role during cardiac development31,36,37), identifying overexpressed genes, computing interaction probabilities and discovering significant interactions based on default parameters. This pipeline was performed using all cell classes present in each region (except for platelet–red blood cells) to calculate potential CCIs.
Developmental trajectory analysis
To identify a developmental trajectory of vCMs within our scRNA-seq dataset, we used the Waddington-OT (v.1.0.8)72 package. The vCM cell class was isolated, which represents subpopulations C1–C11 of the cardiomyocyte compartment, and the corresponding cells were used for Waddington-OT trajectory inference as outlined in GitHub (https://broadinstitute.github.io/wot/tutorial/) without the optional step of estimating initial growth rates. Transport matrices were calculated for each adjacent pair of time points (9 p.c.w.–11 p.c.w., 11 p.c.w.–13 p.c.w., and 13 p.c.w.–15 p.c.w.) and then the trajectory was computed by calculating the descendent distributions at the 9 p.c.w stage. Normalized pseudotime values used for subsequent analyses were calculated by taking the quantile for each cell ranked by raw pseudotime value. Following the construction of the developmental trajectory, the resulting pseudotime for vCMs was utilized to order the GRN and CCIs of the vCMs by determining the expression-weighted pseudotime of each respective transcription factor and receptor or ligand expressed by vCMs as previously described73.
MERFISH processing
For processing MERFISH data, individual RNA molecules were decoded using MERlin (v.0.6.1) as previously described8. Images were aligned across hybridization rounds by maximizing phase cross-correlation on the fiducial bead channel to adjust for drift in the position of the stage from round to round. Background was reduced by applying a high-pass filter, and decoding was then performed per pixel. For each pixel, a vector was constructed of the 22 brightness values from each of the 22 rounds of imaging. These vectors were then L2-normalized, and their Euclidean distances to each of the L2-normalized barcodes from MERFISH codebook were calculated. Pixels were assigned to the gene whose barcode they were closest to, unless the closest distance was greater than 0.512, in which case the pixel was not assigned a gene. Adjacent pixels assigned to the same gene were combined into a single RNA molecule. Molecules were filtered to remove potential false positives by comparing the mean brightness, pixel size and distance to the closest barcode of molecules assigned to blank barcodes versus molecules assigned to genes to achieve an estimated misidentification rate of 5%. The exact position of each molecule was calculated as the median position of all pixels consisting of the molecule.
Cellpose (v.1.0.2)74 was used to perform image segmentation to determine the boundaries of cells and nuclei. Distinct RNA molecules were identified and assigned to individual cells as segmented by total polyadenylated RNA staining and DAPI staining, which allowed for the detection of cellular boundaries. The nuclei boundaries were determined by running Cellpose with the ‘nuclei’ model using default parameters on the DAPI stain channel of the pre-hybridization images. Cytoplasm boundaries were segmented with the ‘cyto’ model and default parameters using the polyT stain channel. RNA molecules identified by MERlin were assigned to cells and nuclei by applying these segmentation masks to the positions of the molecules. Any segmented cells that did not have any barcodes assigned were removed before constructing the cell-by-gene matrix.
smFISH computational analysis
Images were flatfield-corrected for the two gene channels (750 nm and 635 nm) and the fiducial marker (405 nm) channel. To reduce background noise for each hybridization round, the images of the preceding hybridization round were reduced in intensity and subtracted to obtain new background-subtracted images. The images were then locally normalized by subtracting a 15 × 15 blur from each pixel before undergoing maximum intensity projection into two dimensions. For transcript detection, the OpenCV function adaptiveThreshold was used with a block size of 41 pixels, and a subtracted constant ranging from −80 to −70 among our replicate smFISH experiments. This constant was empirically determined by choosing a value that ensured the resulting mask only captured visible fluorescent spots across diverse imaging planes for each gene. Using the regionprops function from Scikit-Image, we filtered out spots with an eccentricity value of 0 and cells with low pixel area (<4 pixels) to combat artefactual fluorescence detection. A global threshold was identified for the images of each gene by evaluating the values of features determined as nonspecific background (for example, irregular shape, low intensity). The coordinates of local brightness maxima that remained unattenuated after applying this global threshold were stored. Coordinates lying within the adaptiveThreshold mask boundaries were identified and counted as a single identified gene transcript. The images for each of the smFISH imaging rounds were aligned to the respective initial MERFISH hybridization round images to correct for microscopic drift using the fiducial marker channel. This was done by fitting spots to the fiducial bead markers of both sets of images, then minimizing the median distances between them. DAPI segmentation masks obtained from the MERFISH imaging were translated using this drift correction so that all identified gene transcript locations could accurately be assigned to the drift-corrected nuclei, which enabled reconstruction of a spatial mosaic of the cellular gene expression for each of our sequentially imaged gene targets. For the replicate smFISH experiments to validate MERFISH gene markers, each gene was imaged twice on the same colour channel but in different non-consecutive rounds, allowing for a more robust analysis by using the combination of the two images to reduce the effect of noise in the result. Genes were imaged on three colour channels: Alexa 750, Cy5 and Cy3. The genes were separated and analysed into batches of six, with the imaging being done in a pattern described as follows. In the first imaging round, genes A, B and C were imaged on the Alexa 750, Cy5 and Cy3 channels, respectively. On the second round, genes D, E and F were imaged on the same three channels. The third and fourth imaging rounds were then repeats of the first and second rounds. By imaging each gene twice, the data could be analysed as a pseudo-MERFISH experiment, whereby a codebook was designed with each gene having a barcode containing two ‘on’ bits in the two imaging rounds they were imaged. Using this codebook, the data was processed using the same method as the MERFISH data as previously described8.
Cell clustering analysis of MERFISH
With the cell-by-gene matrix, we followed a standard procedure as suggested in the Scanpy (v.1.8)75 tutorial using Python (v.3.9) for processing MERFISH data. Count normalization, principal component analysis (PCA), neighbourhood graph construction and UMAP were performed using the default parameters in Scanpy. We performed Leiden clustering utilizing a resolution of 2 because the additional clusters gained at higher resolutions either did not have differentially expressed genes or were related to technical imaging artefacts. The top 20 differential genes identified by the rank_gene_groups() function were used to annotate each cluster. We further subclustered the vCM clusters using Leiden clustering at a default resolution of 1 to further annotate vCMs as compact and trabecular vCMs for both the left and right ventricles. To investigate the cellular populations in the ventricle specifically (both 13 p.c.w. and 15 p.c.w.), we manually defined the ventricular region, subsetted the MERFISH dataset to those cells populating the ventricle and performed Leiden clustering using a similar strategy to that used in the overall clustering (resolution of 5). Again, the top 20 genes identified by the rank_gene_groups() function were used to annotate each cluster.
Integration of the scRNAseq and MERFISH datasets
To integrate the scRNA-seq and MERFISH datasets, we first isolated both datasets to only the 238 MERFISH target genes interrogated by both modalities. We then utilized Scanpy’s implementation of Harmony to project both the scRNA-seq and MERFISH datasets into a shared PCA space76. The dimensionality of the joint embedding was visualized using UMAP (min_dist=0.3, 30 nearest neighbours in Pearson correlation distance). The parameters matched those used in a previous publication of Harmony76. To impute a complete expression profile and cell label for each MERFISH profile, we assigned the expression profile and cell label of the closest scRNA-seq cell to the MERFISH cell in the Harmony PCA space using the Euclidean distance metric (default number of PCs).
To evaluate the performance of the gene imputation method, we developed a predictability score for each gene which is the Pearson correlation between the imputed expression and measured image expression for all genes (Supplementary Fig. 14a). Because the shared embedding space is constructed using the 238 MERFISH target genes, it is expected that these genes would have higher predictability scores than genes not used in the construction. To avoid this bias, tenfold cross-validation was used to calculate independently the MERFISH and scRNA-seq gene predictability scores. To this end, a new shared embedding utilizing only 90% of the 238 MERFISH target genes was used to calculate the MERFISH and scRNA-seq gene predictability scores for the remaining 10% of genes that were not included for constructing the embedding. This process was repeated 10 times with a different 10% of genes being imputed by a different shared embedding each time to cover the full set of 238 genes. To calculate whether a predictability score represented a prediction that is a significant improvement over random prediction, we calculated predictability scores using a randomly connected neighbour graph. In other words, rather than predicting the expression from the cell with the most similar gene expression, the prediction was made from a randomly selected cell in the dataset, and then the predictability score was calculated between the measured expression values and these randomly predicted values. This process was repeated 100 times to estimate a normal distribution of predictability scores that result from random prediction. P values were then determined for the true predictability scores based on rejecting the null hypothesis that the true scores originated from the null normal distributions. Finally, these P values were corrected for multiple hypothesis testing using the Bonferroni method. We observed that the maximum scRNA-seq predictability score for a gene that failed this significance test (adjusted P value > 0.01) was 0.11.
Identifying CCs
We sought to define CCs that represented distinct shared cellular environments defined by the neighbouring cells in close proximity. To this end, we clustered each MERFISH cell based on the cell composition of neighbouring cells within a 150 μm zone, which represented a typical diffusion distance for extracellular signalling molecules77. This zone sampled approximately 300 neighbours. The zone of each cell was therefore represented by a vector containing the relative proportions of each of the 27 identified cells in both the overall and ventricular subset of the MERFISH dataset. We then clustered the zones using Python’s scikit-learn (v.0.22) implementation of Kmeans with k = 13 for the overall MERFISH dataset or k = 9 for the ventricular subset of the MERFISH dataset, chosen by silhouette score. Thus, each MERFISH cell was assigned to 1 of the 13 or 9 CCs in the overall or ventricular subset of the MERFISH dataset, respectively.
To infer community-specific CCIs, cell annotations derived for each MERFISH cell were transferred to the nearest scRNA-seq profile in the Harmony joint embedding space and used for the pipeline of CellChat as described in the ‘CCI analysis’ section. The overall and ventricular communities were each analysed separately by analysing the scRNA-seq profiles assigned to those communities. For each overall or ventricular cellular community, we only considered cells that represented at least 5% or 3.5% of the community in the MERFISH dataset for CellChat CCI analysis.
Connectivity, ventricular wall depth and pseudotime analyses of vCMs
To visualize the similarity in the gene expression profiles of the vCMs, we isolated the vCM-LV-compact I, vCM-LV-compact II, vCM-LV-hybrid, vCM-LV-trabecular I, vCM-LV-trabecular II and vCM-LV/RV-Purkinje populations, and reperformed PCA, nearest neighbour graph construction and UMAP utilizing Scanpy with default parameters. We then used Scanpy’s implementation of partition based graph abstraction to construct a graph in which the nodes represent different vCMs, and the edges represent the degree of connectivity between the vCMs in the neighbourhood graph. This captured a measure of similarity between the vCMs.
To determine the distance of each MERFISH cell within the ventricular wall, the ventricular wall depth of each MERFISH cell was defined as the distance to the nearest epicardial cell or EPDC, which both lie on the outer layer of the heart. To compare ventricular wall depth between different ventricles, the wall depth values were normalized by dividing each value by the maximum depth of the corresponding ventricle. To identify depth correlated genes, we computed the Pearson correlation coefficient between expression and ventricular wall depth for each gene. We considered genes with a correlation greater than 0.2 to be depth correlated. Next, the diffusion pseudotime distance on the vCM nearest neighbour graph from the medoid of the vCM-LV-compact I cluster were calculated using Scanpy’s scanpy.tl.dpt() function with default parameters. We note that this metric is often reported as pseudotime to represent developmental changes, but in this case, we use it simply as a metric for expression similarity to vCM-LV-compact I cells. The scaled expression of the top genes correlating with pseudotime were plotted as a smoothed spline (Extended Data Fig. 6d) as previously described78.
Migration distance measurement
The migration distance of the bioprinted hPSC-CMs was measured by using the D0 position as the starting point and calculating the distance migrated by the hPSC-CMs daily. In brief, brightfield and fluorescent (green and red) confocal images of the GFP+ and NLS-mKATE2+ hPSC-CMs were taken on a Leica SP5, with the brightfield images used to visualize the construct. Because of minor variations in size and cell number between printed constructs, we normalized the migration distances by first dividing the width of the construct into 15 even blocks for each image. Within each block, the distances from the D0 position to the furthest position (for each day) of the hPSC-CMs were calculated. We then averaged the distances measured for the 15 blocks to calculate the migration distance of each condition and line.
Statistics and reproducibility
Replicates and statistical tests are described in the figure legends. Sample sizes were not predetermined utilizing statistical methods. Tissue samples were not randomized, nor were the investigators blinded during collection as no subjective measurements were taken. Data for scRNA-seq and MERFISH were collected from all available samples and no randomization was necessary. For the studies utilizing human pluripotent stem cell lines, treatment with NRG1 was randomly assigned. For the animal studies, animals were randomly chosen from each genotype and stage. Blinding during analysis was not necessary as all of the results were analysed with the use of unbiased analysis and software tools that are not affected by the sample. All experiments were independently repeated at least three times with similar results, including experiments in Fig. 5e, Extended Data Figs. 2a and 12a and Supplementary Fig. 18a. To identify differentially expressed genes between clusters, a Wilcoxon rank-sum test was performed and the resulting P value was corrected using the Bonferroni procedure. For the scRNA-seq predictability scores, the P values were generated by using bootstrapping to generate a distribution of scores for each gene and then calculating (1– cumulative distribution function) to obtain the P value. For the migration distance, ventricular wall thickness and qPCR results, we used a one-way analysis of variance using R (v.4.2.0; https://www.r-project.org/).
The sample sizes for the violin plot in Fig. 2e are listed as follows: from top to bottom, n = 9,106, 7,661, 19,901, 3,791, 4,003, 60,810, 28,263, 16,369, 6,956, 21,087, 17,940, 5,135 and 27,613 cells. Fig. 4c: from left to right, n = 541, 849, 1,552, 719, 2,290, 1,112, 754, 499, 335, 55, 13, 701, 338, 177, 163 and 49 cells. Extended Data Fig. 5d: from left to right, n = 9,106, 7,661, 19,901, 3,791, 4,003, 60,810, 28,263, 16,369, 6,956, 21,087, 17,940, 5,135 and 27,613 cells. Extended Data Fig. 5e: from left to right, n = 27,613, 5,135, 17,940, 21,087, 7,661, 6,956, 3,791, 28,263, 9,106, 4,003, 60,810, 16,369 and 19,901 cells. Extended Data Fig. 8f: top panel from left to right (13 p.c.w./15 p.c.w.), n = 573/706, 976/160, 1,532/895, 354/303, 784/553, 720/NA, 187/NA, 1,440/866, 1,905/840 and 508/417 cells; bottom panel from left to right (13 p.c.w./15 p.c.w.), n = 711/274, 313/305, 548/711, 387/21, 1,444/938, 800/451, 557/409, 79/80 and 66/124 cells. Extended Data Fig. 9d: from left to right, n = 9,723, 18,908, 21,203, 8,042, 47,906, 16,225, 5,814, 6,307 and 18,592 cells. Extended Data Fig. 9e: from left to right, n = 18,592, 8,042, 5,814, 6,307, 47,906, 21,203, 18,908, 16,225 and 9,723. Extended Data Fig. 11e: from left to right, n = 81,880, 75,531, 34,953, 145,935, 19,949 and 18,485 RNA molecules. Violin plots consisting of cell numbers of ten or fewer are not shown and are labelled as ‘NA’ in those cases.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.